Publications
Our research sits at the interface of genomics, RNA biology, and machine learning, with a particular focus on understanding how sequence and chromatin architecture shape gene regulation across cells, tissues, and disease states. Across these publications, we develop computational methods and foundation models for predicting 3D genome organization and gene expression, design and interpret models of RNA splicing, build tools for long-read epigenetic analysis, and use large-scale transcriptomic and genetic data to uncover mechanisms of cancer, hematopoiesis, neurodegeneration, and other complex traits. Together, this work reflects a broad effort to turn high-dimensional genomic data into interpretable models of regulatory biology.
Highlighted
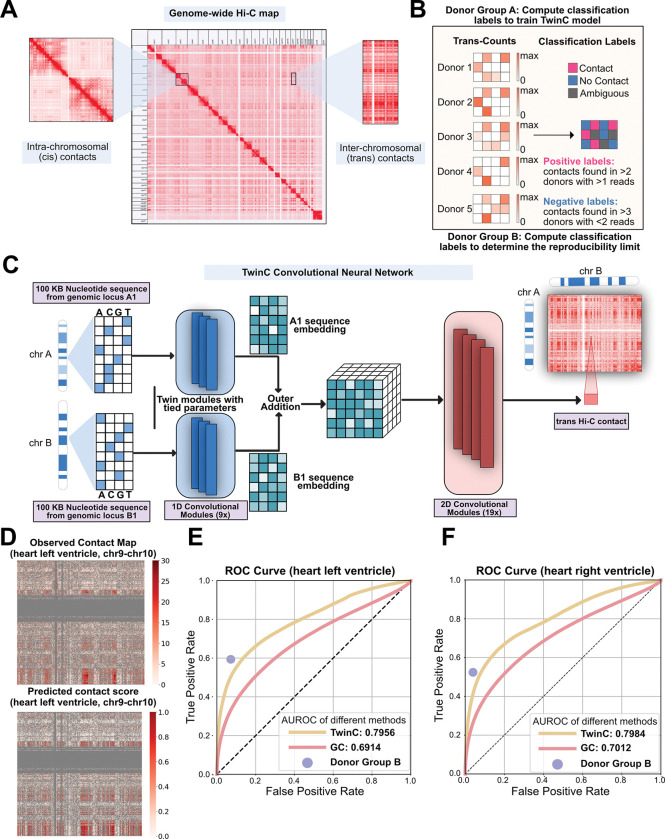
Prediction and functional interpretation of inter-chromosomal genome architecture from DNA sequence with TwinC
openRxiv
·
21 Sep 2024
·
doi:10.1101/2024.09.16.613355
Predicts and interprets inter-chromosomal genome architecture directly from DNA sequence.
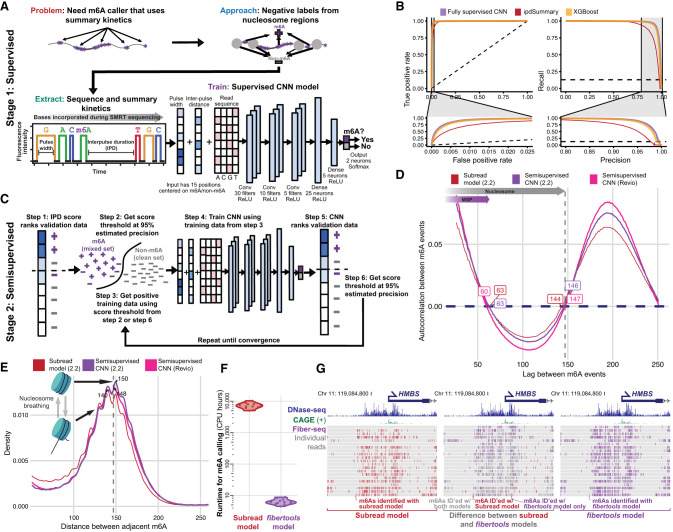
DNA-m6A calling and integrated long-read epigenetic and genetic analysis with fibertools
Genome Research
·
07 Jun 2024
·
doi:10.1101/gr.279095.124
Describes DNA-m6A calling and integrated long-read epigenetic analysis with fibertools.
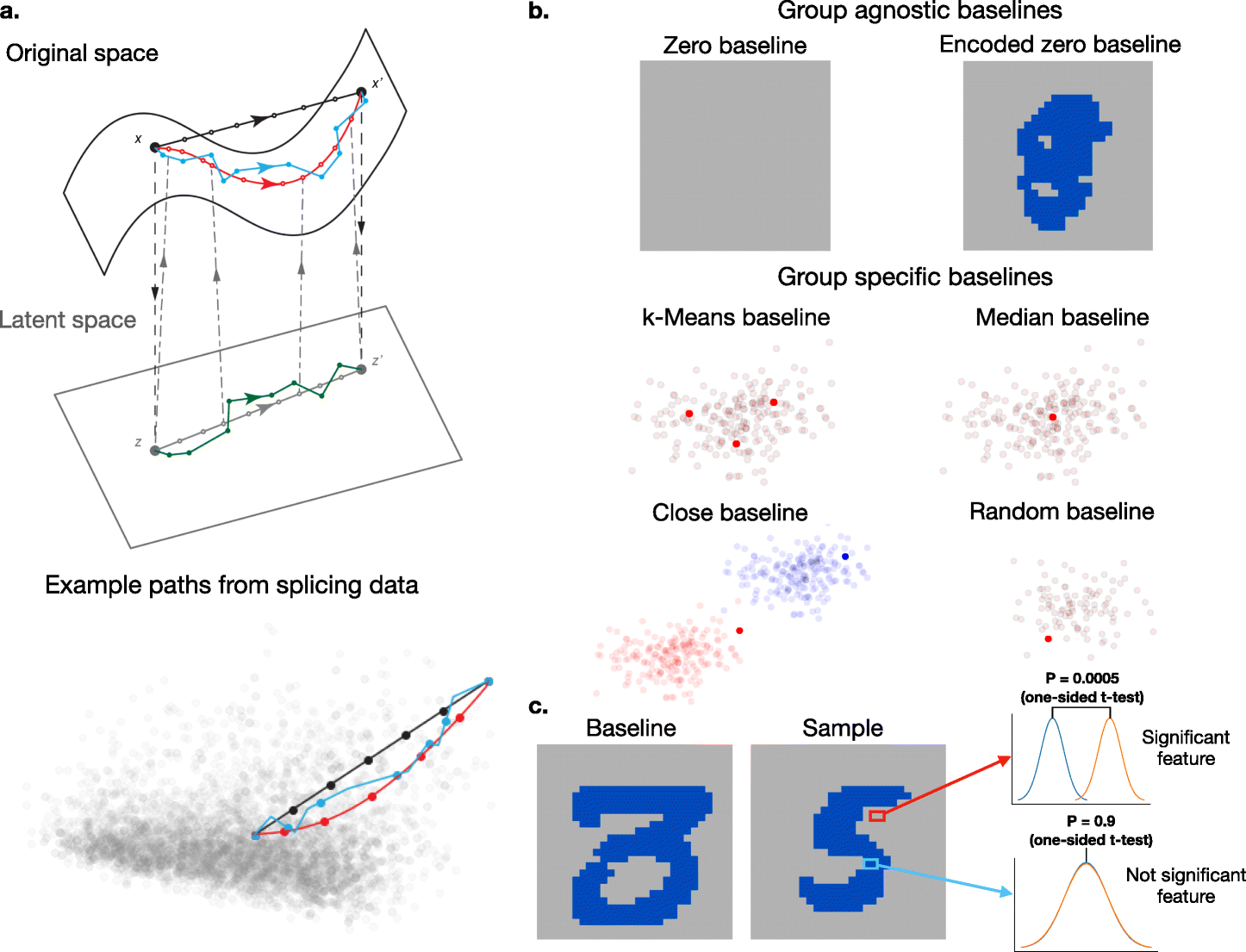
Enhanced integrated gradients: improving interpretability of deep learning models using splicing codes as a case study
Genome Biology
·
19 Jun 2020
·
doi:10.1186/s13059-020-02055-7
Improves deep learning interpretability with Enhanced Integrated Gradients using splicing codes as a case study.
All
2025
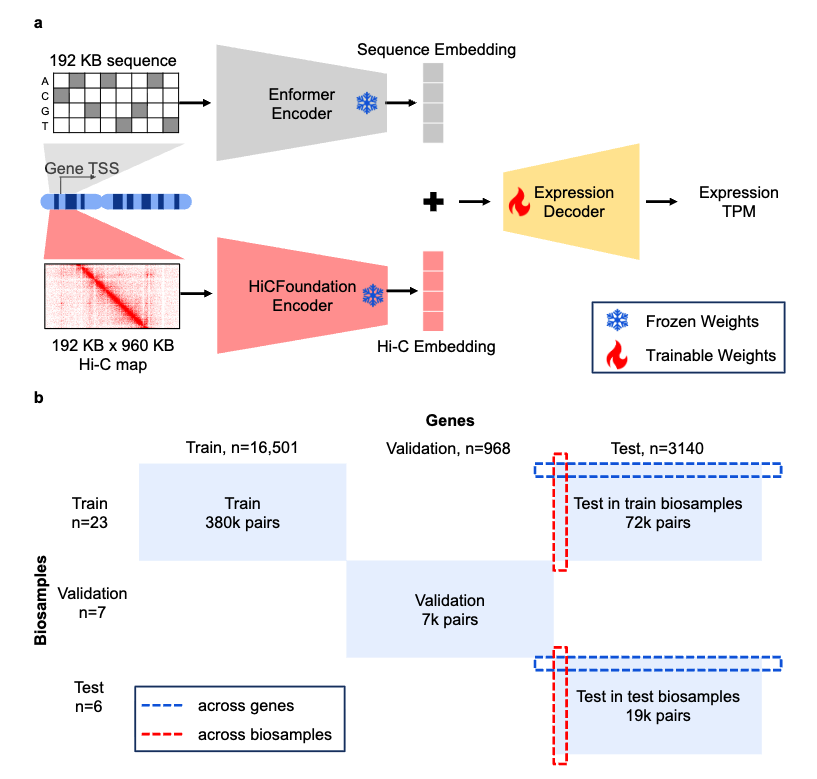
Puget predicts gene expression across cell types using sequence and 3D chromatin organization data
bioRxiv
·
20 Nov 2025
·
doi:10.1101/2025.11.19.689320
Predicts gene expression across cell types by combining sequence information with 3D chromatin organization.
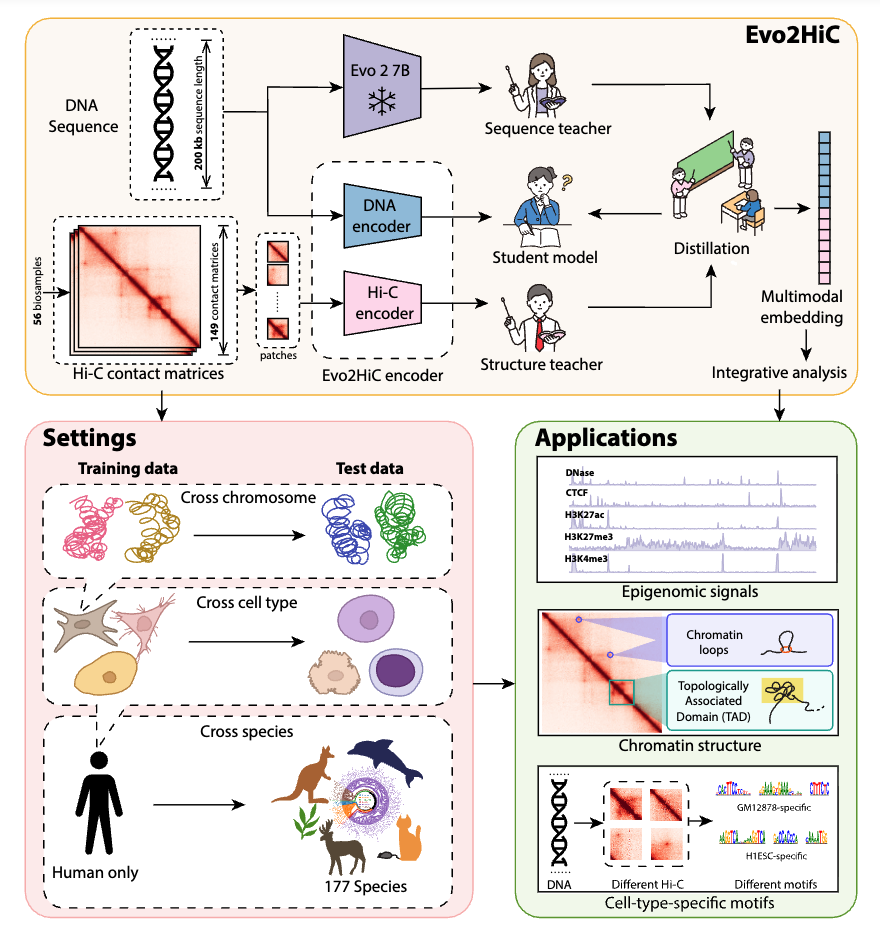
Evo2HiC: a multimodal foundation model for integrative analysis of genome sequence and architecture
bioRxiv
·
19 Nov 2025
·
doi:10.1101/2025.11.18.689171
Introduces a multimodal foundation model that jointly learns genome sequence and chromatin architecture.
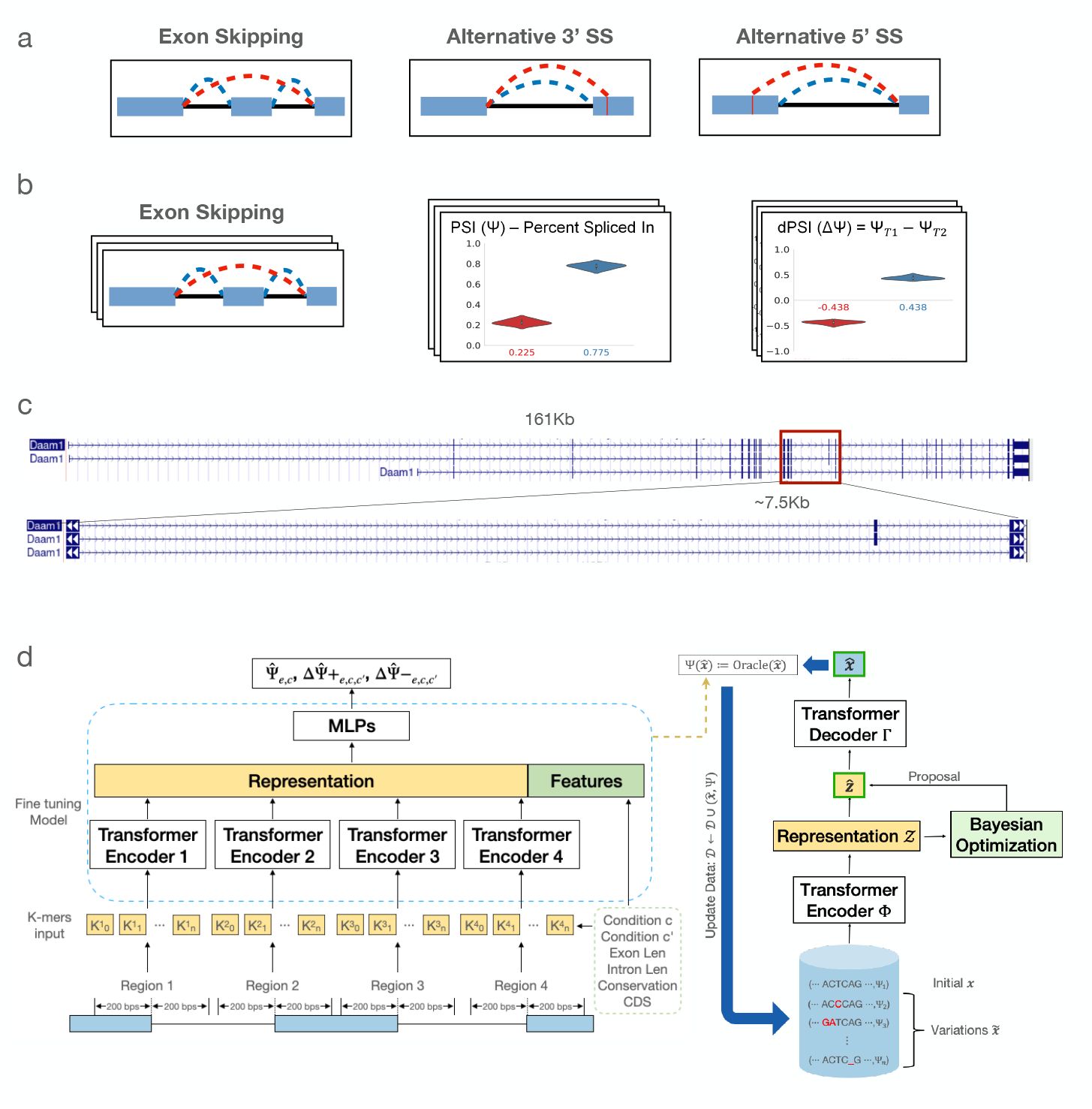
Generative modeling for RNA splicing prediction and design
openRxiv
·
24 Jan 2025
·
doi:10.1101/2025.01.20.633986
Uses generative modeling to predict and design RNA splicing outcomes from sequence features.
2024
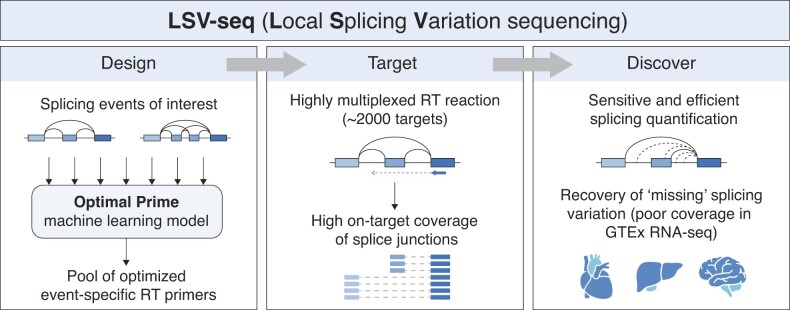
Machine learning-optimized targeted detection of alternative splicing
Nucleic Acids Research
·
27 Dec 2024
·
doi:10.1093/nar/gkae1260
Applies machine learning to optimize targeted detection of alternative splicing events.

A generalizable Hi-C foundation model for chromatin architecture, single-cell and multi-omics analysis across species
openRxiv
·
20 Dec 2024
·
doi:10.1101/2024.12.16.628821
Presents a generalizable Hi-C foundation model for chromatin architecture across species and assays.

Prediction and functional interpretation of inter-chromosomal genome architecture from DNA sequence with TwinC
openRxiv
·
21 Sep 2024
·
doi:10.1101/2024.09.16.613355
Predicts and interprets inter-chromosomal genome architecture directly from DNA sequence.
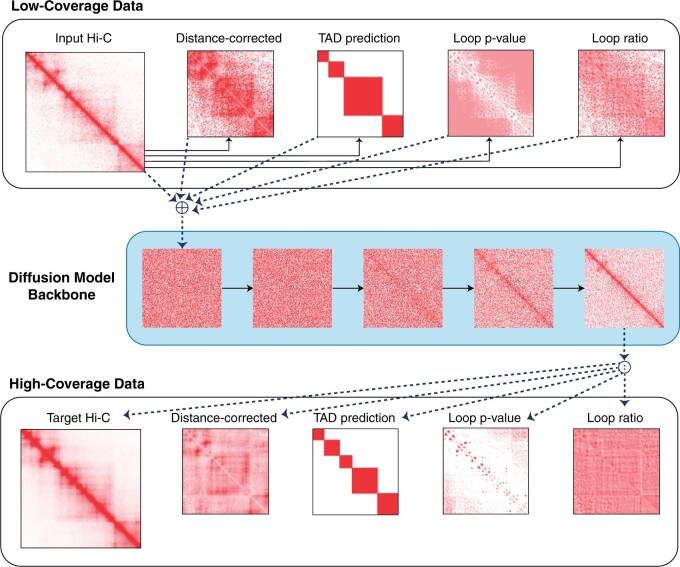
Enhancing Hi-C contact matrices for loop detection with Capricorn: a multiview diffusion model
Bioinformatics
·
28 Jun 2024
·
doi:10.1093/bioinformatics/btae211
Enhances Hi-C contact matrices with a diffusion model to improve loop detection.

DNA-m6A calling and integrated long-read epigenetic and genetic analysis with fibertools
Genome Research
·
07 Jun 2024
·
doi:10.1101/gr.279095.124
Describes DNA-m6A calling and integrated long-read epigenetic analysis with fibertools.
2023
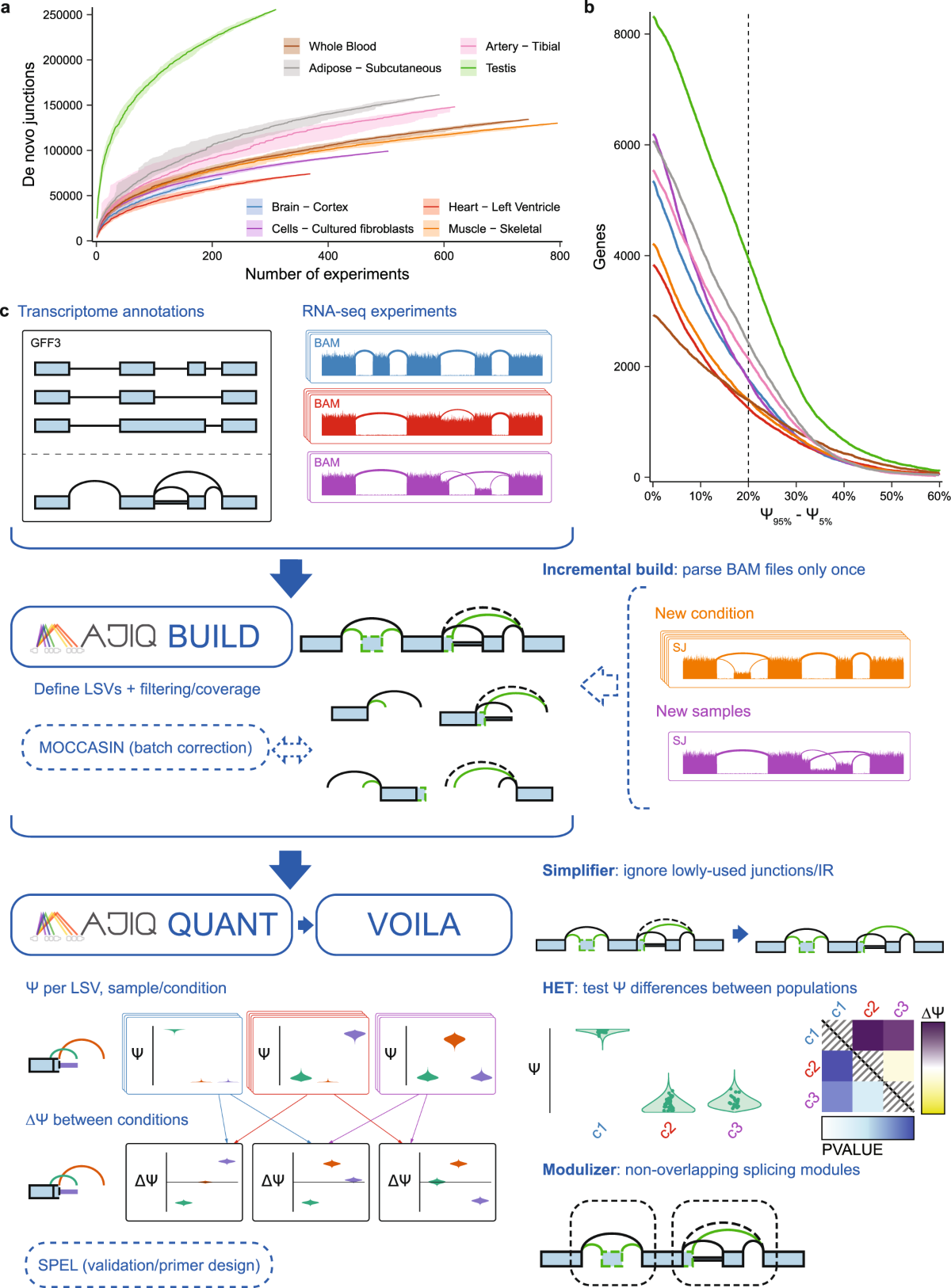
RNA splicing analysis using heterogeneous and large RNA-seq datasets
Nature Communications
·
03 Mar 2023
·
doi:10.1038/s41467-023-36585-y
Studies large heterogeneous RNA-seq datasets to improve RNA splicing analysis at scale.
2022
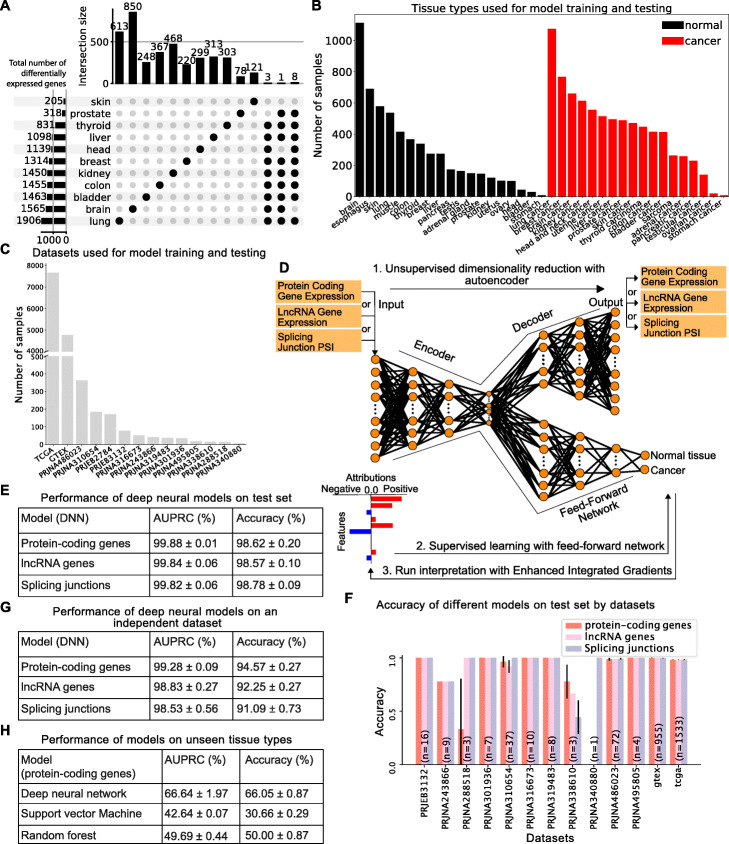
Identifying common transcriptome signatures of cancer by interpreting deep learning models
Genome Biology
·
17 May 2022
·
doi:10.1186/s13059-022-02681-3
Interprets deep learning models to identify transcriptomic signatures shared across cancers.
2021
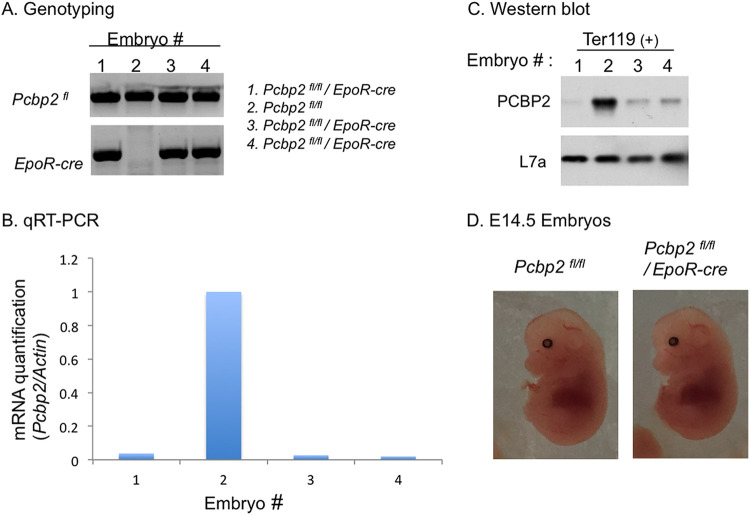
RNA-binding proteins PCBP1 and PCBP2 are critical determinants of murine erythropoiesis
Molecular and Cellular Biology
·
24 Aug 2021
·
doi:10.1128/MCB.00668-20
Shows that PCBP1 and PCBP2 are critical regulators of murine erythropoiesis.
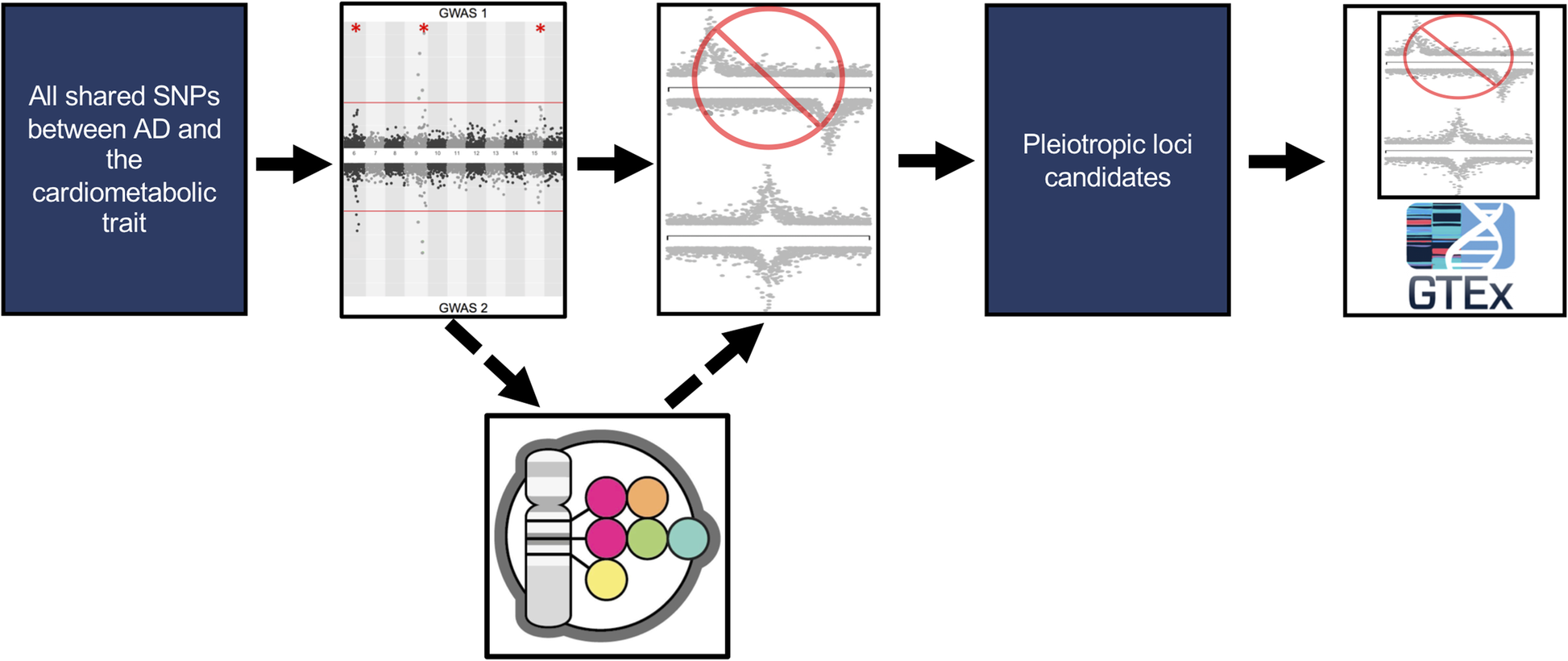
Multi-trait association studies discover pleiotropic loci between Alzheimer’s disease and cardiometabolic traits
Alzheimer's Research & Therapy
·
04 Feb 2021
·
doi:10.1186/s13195-021-00773-z
Identifies pleiotropic loci shared between Alzheimer’s disease and cardiometabolic traits.
2020

Enhanced integrated gradients: improving interpretability of deep learning models using splicing codes as a case study
Genome Biology
·
19 Jun 2020
·
doi:10.1186/s13059-020-02055-7
Improves deep learning interpretability with Enhanced Integrated Gradients using splicing codes as a case study.
2017
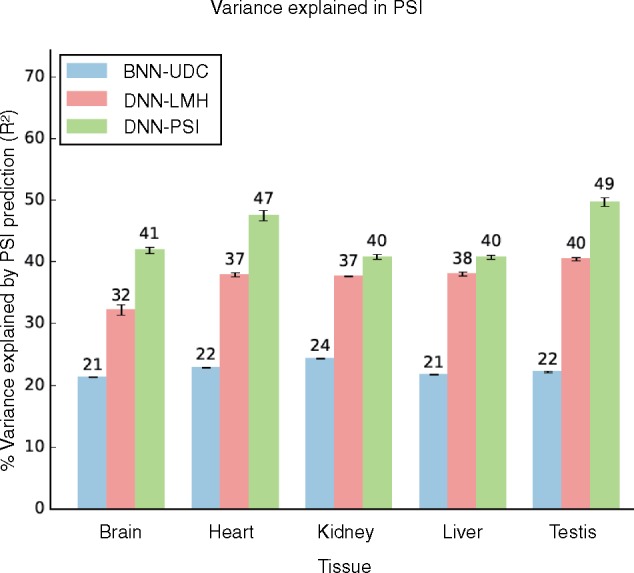
Integrative deep models for alternative splicing
Bioinformatics
·
12 Jul 2017
·
pmc:PMC5870723
Develops integrative deep learning models for predicting alternative splicing regulation.
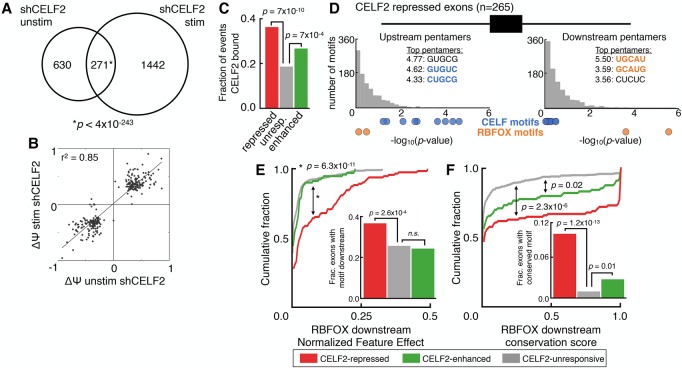
Ancient antagonism between CELF and RBFOX families tunes mRNA splicing outcomes
Genome Research
·
16 May 2017
·
doi:10.1101/gr.220517.117
Reveals how CELF and RBFOX family antagonism shapes mRNA splicing outcomes.
2011

An Optimizing Compiler for Turing Machine Description Language
IUP Journal of Computer Sciences
·
01 Jul 2011
·
iup:tmdl-compiler
Describes an optimizing compiler for a Turing Machine Description Language.