Publications
Highlighted
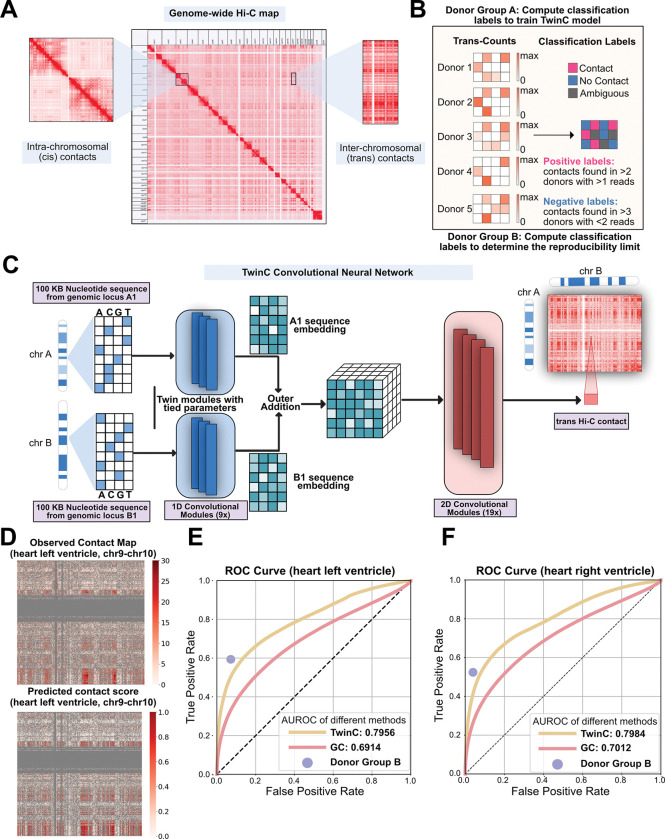
Prediction and functional interpretation of inter-chromosomal genome architecture from DNA sequence with TwinC
openRxiv
·
21 Sep 2024
·
doi:10.1101/2024.09.16.613355
Predicts and interprets inter-chromosomal genome architecture directly from DNA sequence.
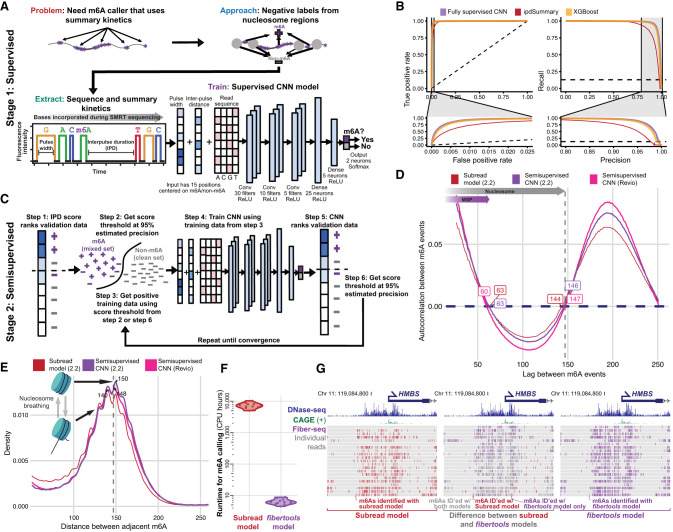
DNA-m6A calling and integrated long-read epigenetic and genetic analysis with fibertools
Genome Research
·
07 Jun 2024
·
doi:10.1101/gr.279095.124
Describes DNA-m6A calling and integrated long-read epigenetic analysis with fibertools.
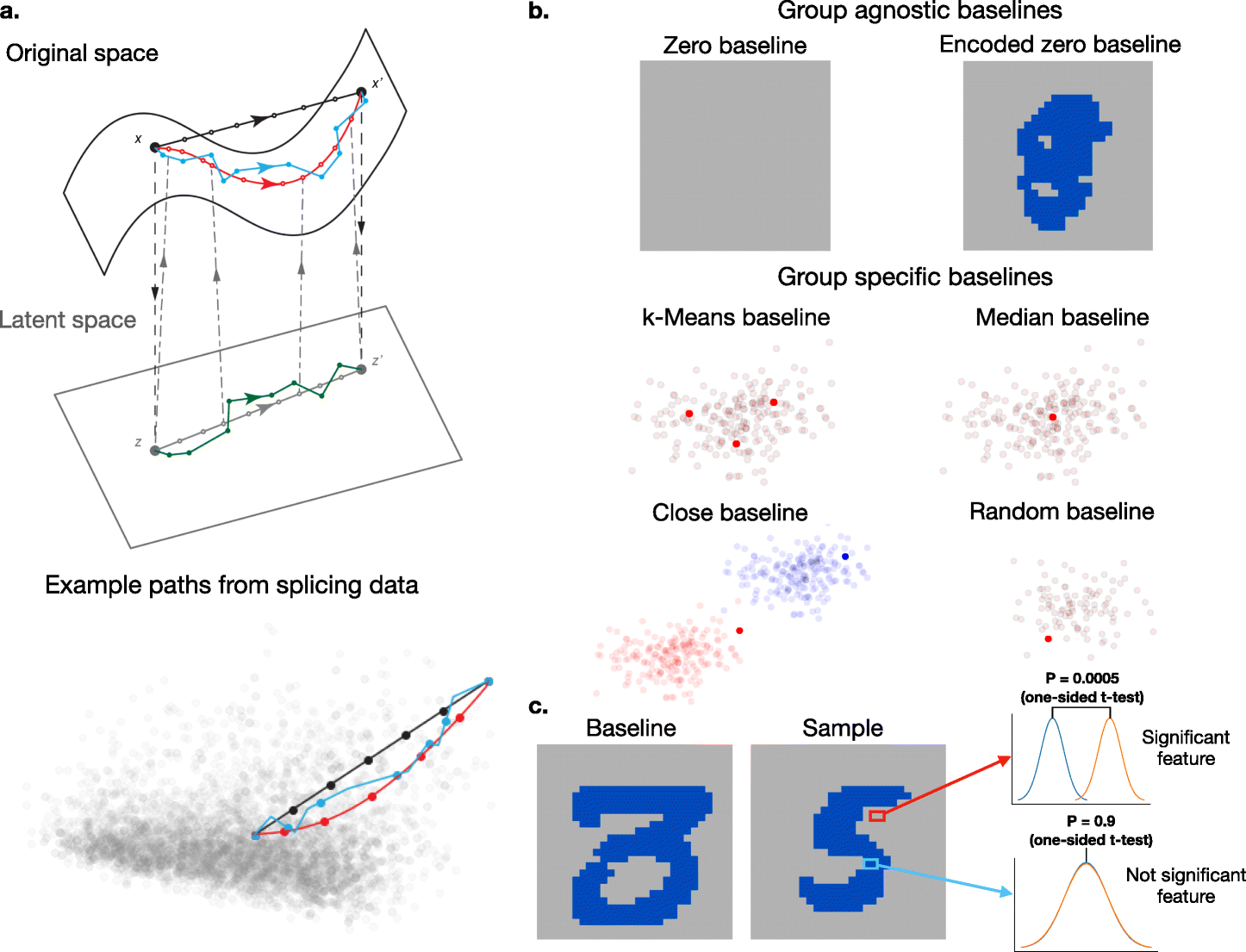
Enhanced integrated gradients: improving interpretability of deep learning models using splicing codes as a case study
Genome Biology
·
19 Jun 2020
·
doi:10.1186/s13059-020-02055-7
Improves deep learning interpretability with Enhanced Integrated Gradients using splicing codes as a case study.
All
2025
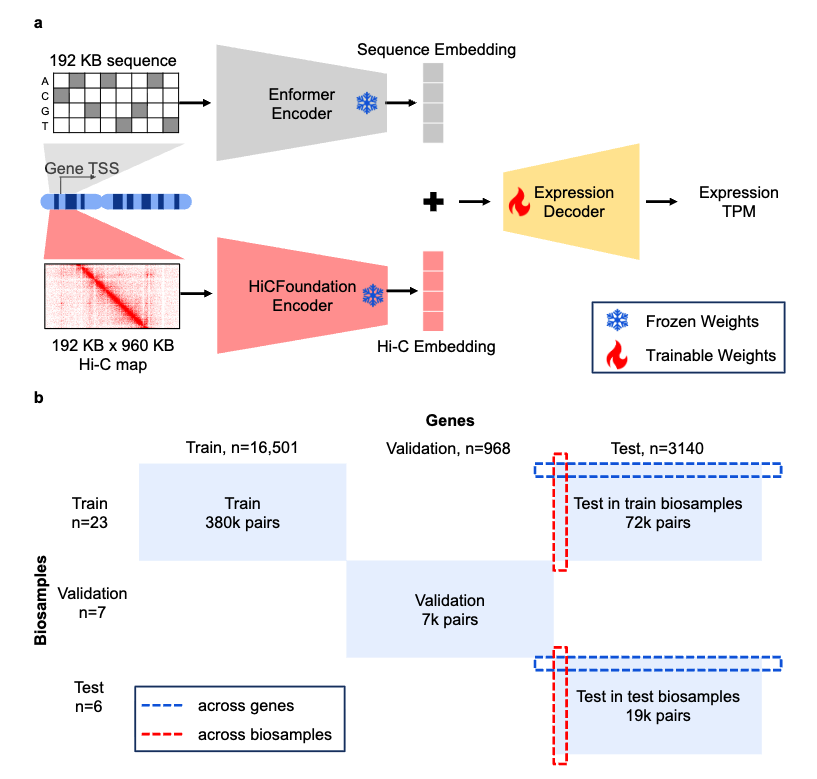
Puget predicts gene expression across cell types using sequence and 3D chromatin organization data
bioRxiv
·
20 Nov 2025
·
doi:10.1101/2025.11.19.689320
Predicts gene expression across cell types by combining sequence information with 3D chromatin organization.
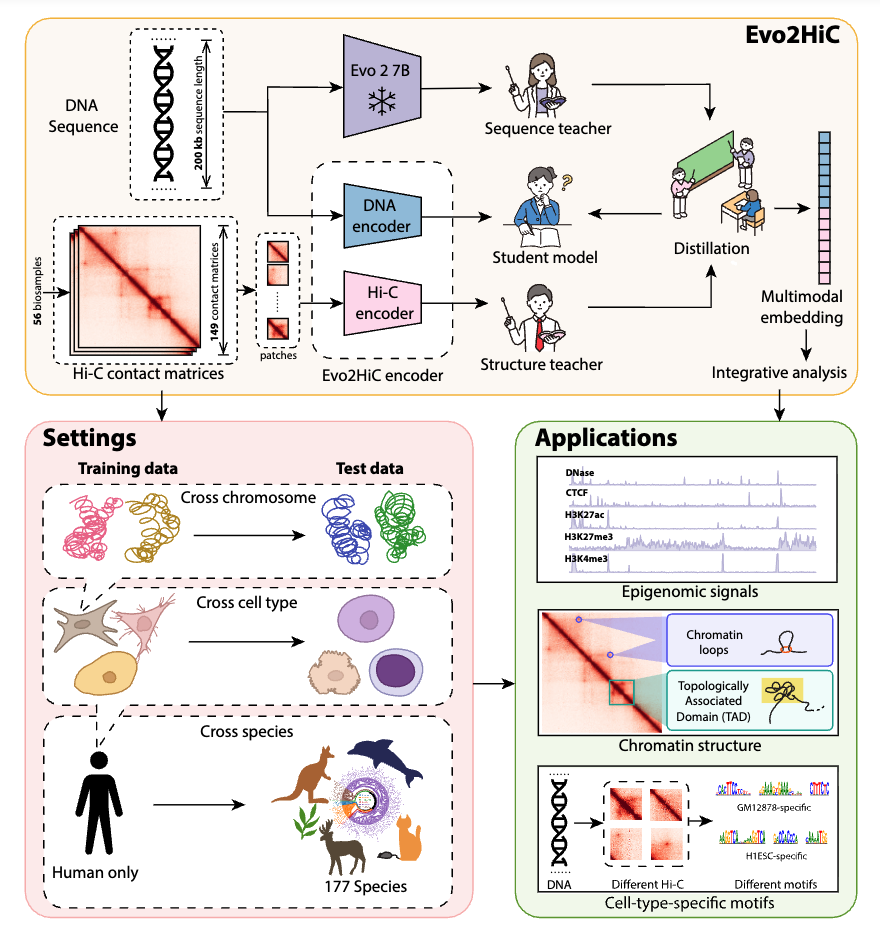
Evo2HiC: a multimodal foundation model for integrative analysis of genome sequence and architecture
bioRxiv
·
19 Nov 2025
·
doi:10.1101/2025.11.18.689171
Introduces a multimodal foundation model that jointly learns genome sequence and chromatin architecture.
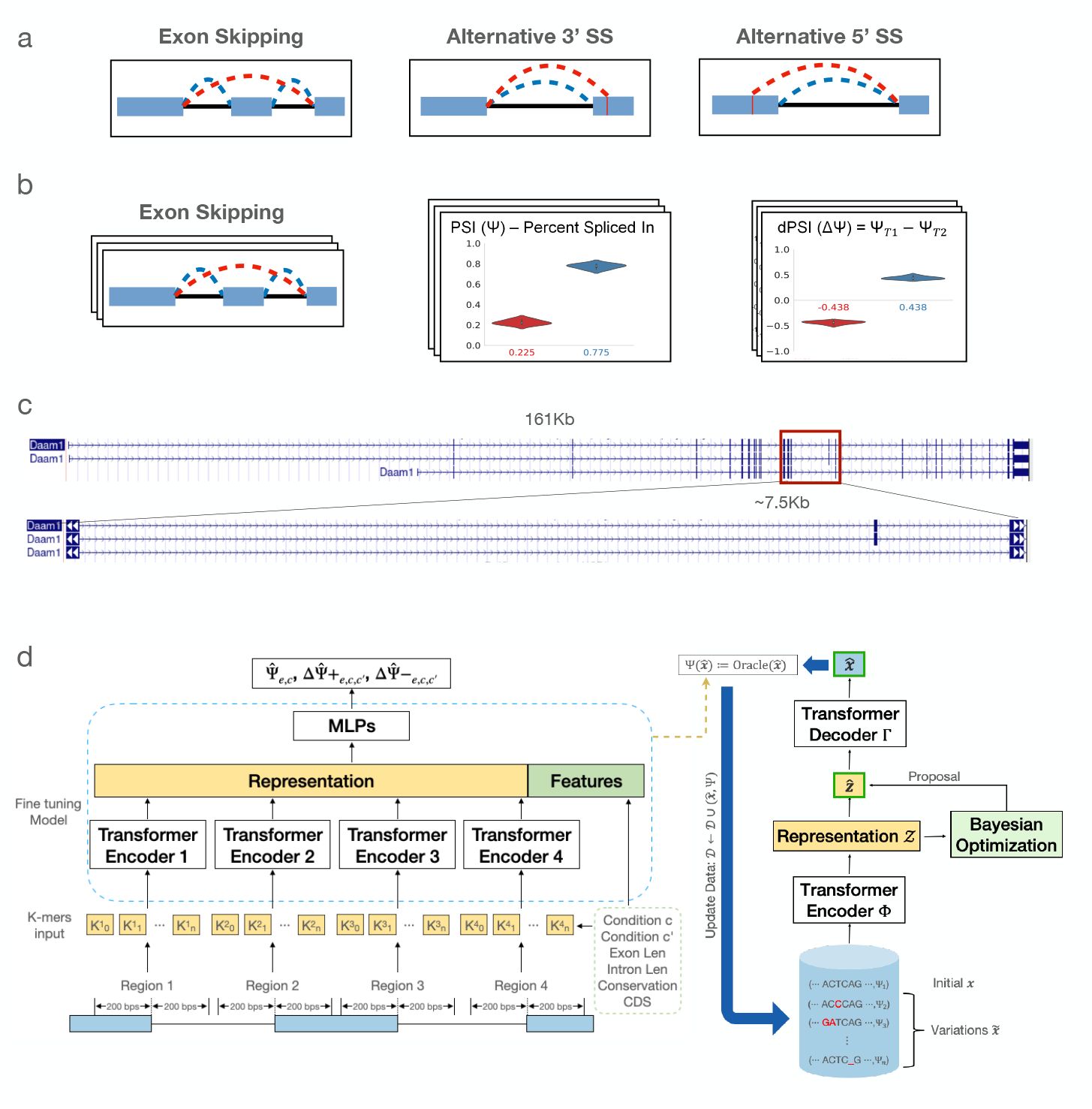
Generative modeling for RNA splicing prediction and design
openRxiv
·
24 Jan 2025
·
doi:10.1101/2025.01.20.633986
Uses generative modeling to predict and design RNA splicing outcomes from sequence features.
2024
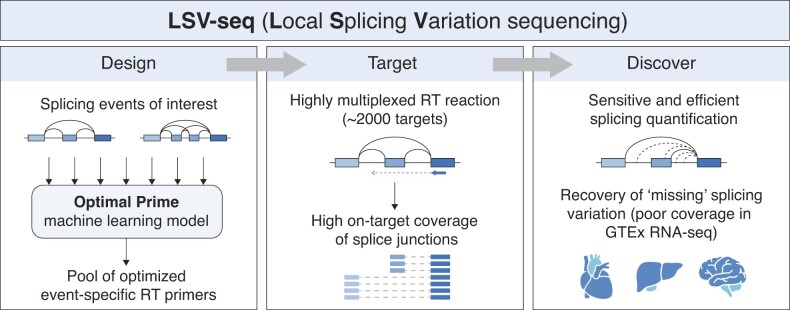
Machine learning-optimized targeted detection of alternative splicing
Nucleic Acids Research
·
27 Dec 2024
·
doi:10.1093/nar/gkae1260
Applies machine learning to optimize targeted detection of alternative splicing events.

A generalizable Hi-C foundation model for chromatin architecture, single-cell and multi-omics analysis across species
openRxiv
·
20 Dec 2024
·
doi:10.1101/2024.12.16.628821
Presents a generalizable Hi-C foundation model for chromatin architecture across species and assays.

Prediction and functional interpretation of inter-chromosomal genome architecture from DNA sequence with TwinC
openRxiv
·
21 Sep 2024
·
doi:10.1101/2024.09.16.613355
Predicts and interprets inter-chromosomal genome architecture directly from DNA sequence.
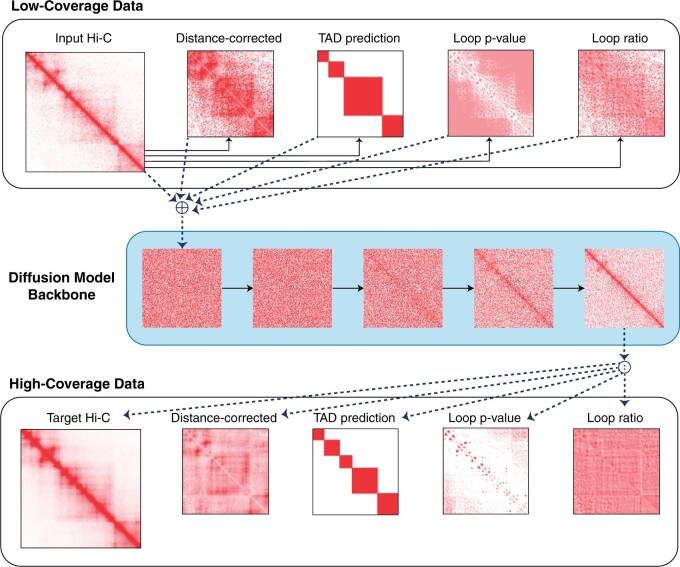
Enhancing Hi-C contact matrices for loop detection with Capricorn: a multiview diffusion model
Bioinformatics
·
28 Jun 2024
·
doi:10.1093/bioinformatics/btae211
Enhances Hi-C contact matrices with a diffusion model to improve loop detection.

DNA-m6A calling and integrated long-read epigenetic and genetic analysis with fibertools
Genome Research
·
07 Jun 2024
·
doi:10.1101/gr.279095.124
Describes DNA-m6A calling and integrated long-read epigenetic analysis with fibertools.
2023
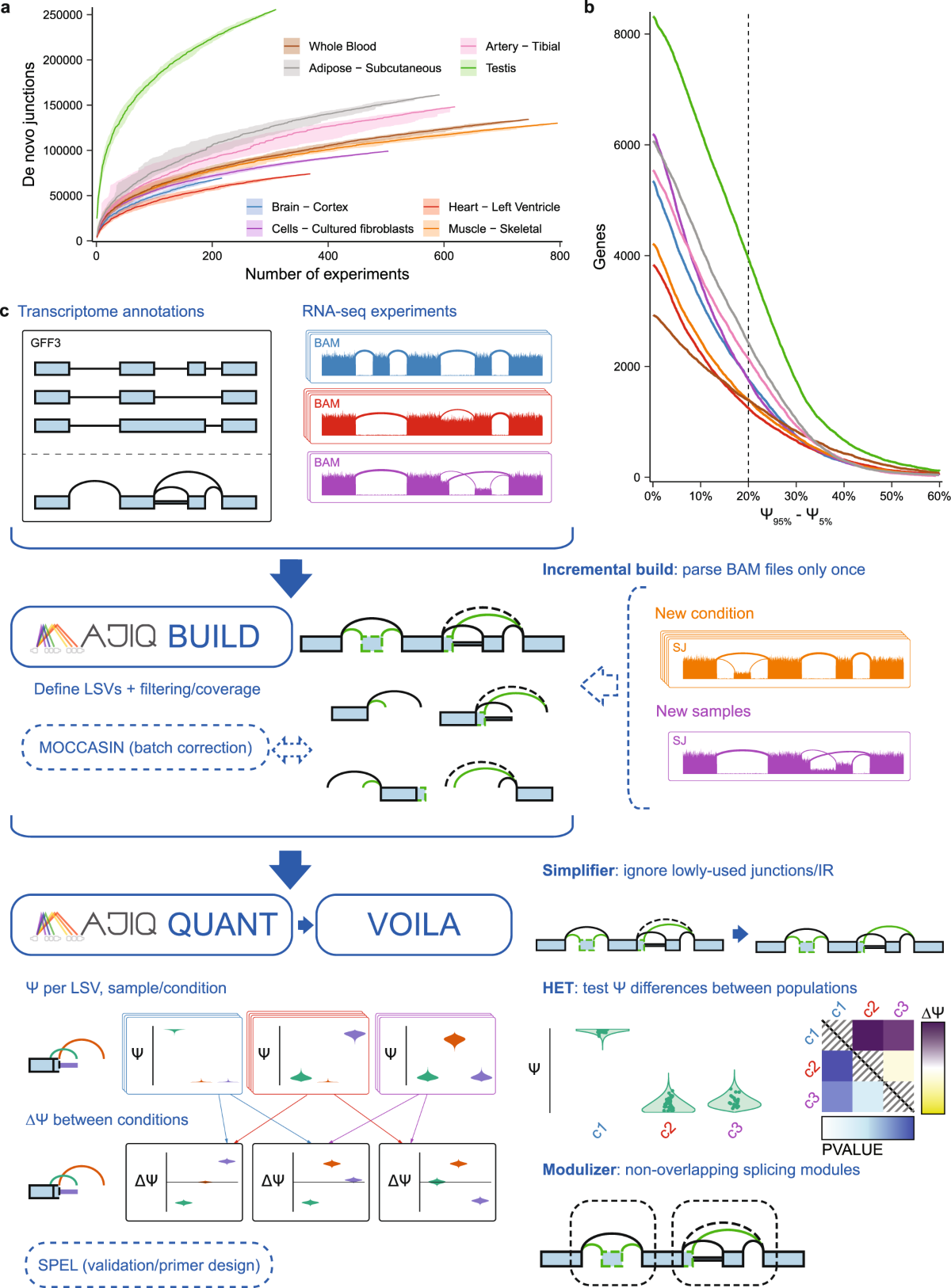
RNA splicing analysis using heterogeneous and large RNA-seq datasets
Nature Communications
·
03 Mar 2023
·
doi:10.1038/s41467-023-36585-y
Studies large heterogeneous RNA-seq datasets to improve RNA splicing analysis at scale.
2022
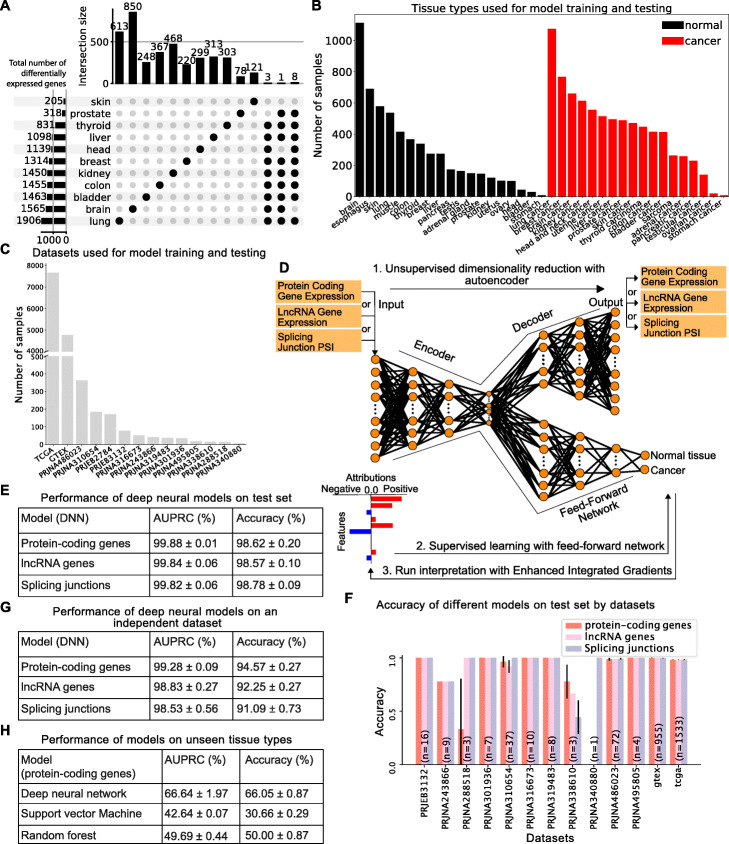
Identifying common transcriptome signatures of cancer by interpreting deep learning models
Genome Biology
·
17 May 2022
·
doi:10.1186/s13059-022-02681-3
Interprets deep learning models to identify transcriptomic signatures shared across cancers.
2021
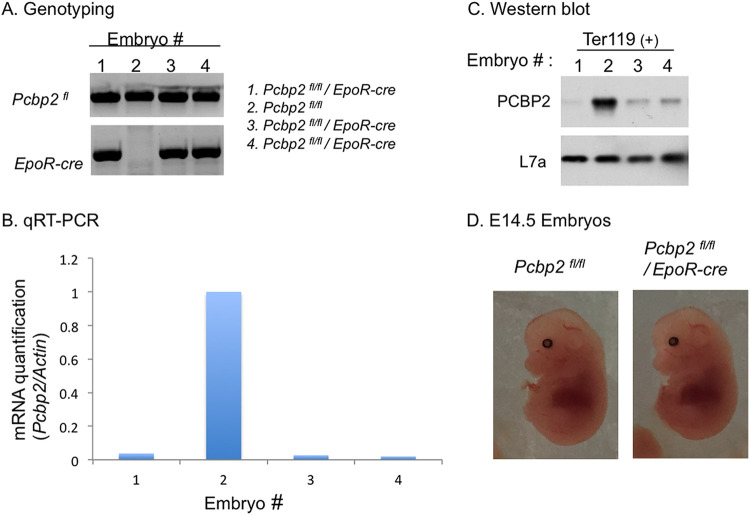
RNA-binding proteins PCBP1 and PCBP2 are critical determinants of murine erythropoiesis
Molecular and Cellular Biology
·
24 Aug 2021
·
doi:10.1128/MCB.00668-20
Shows that PCBP1 and PCBP2 are critical regulators of murine erythropoiesis.
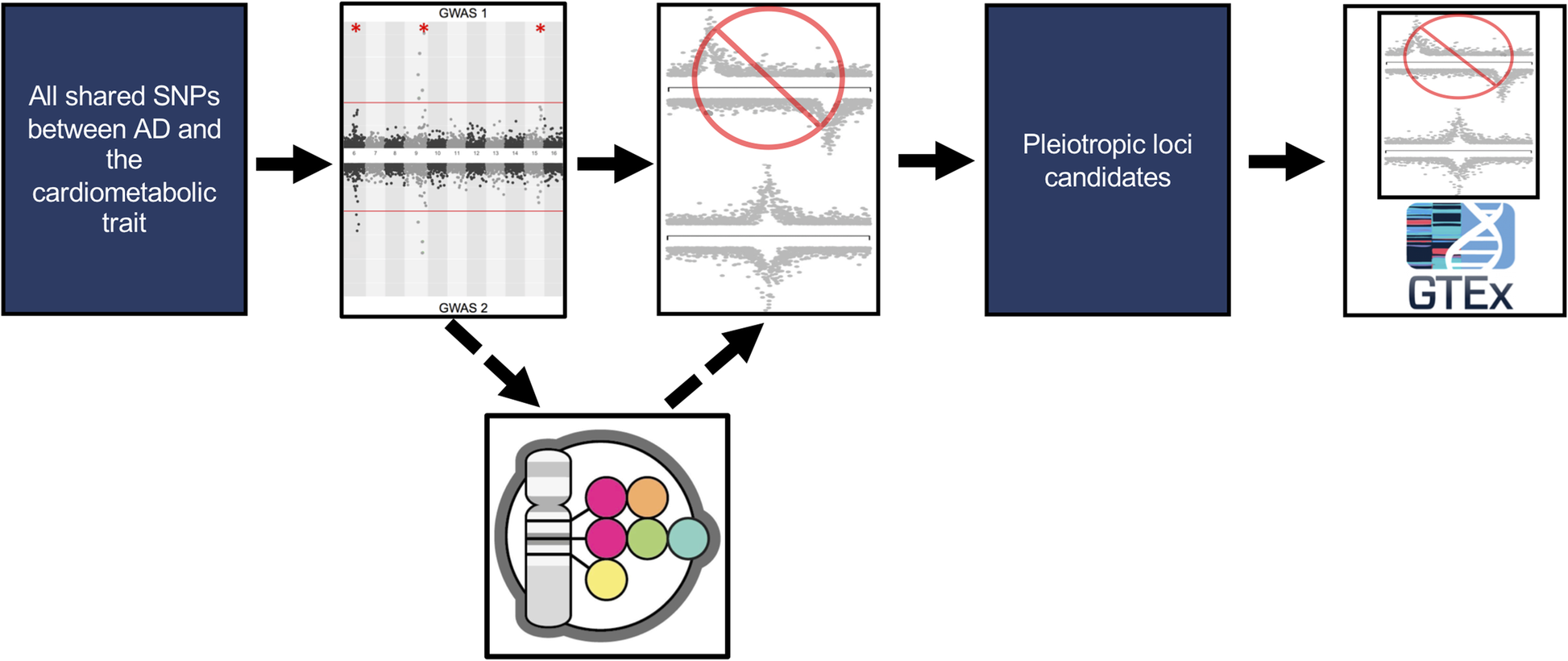
Multi-trait association studies discover pleiotropic loci between Alzheimer’s disease and cardiometabolic traits
Alzheimer's Research & Therapy
·
04 Feb 2021
·
doi:10.1186/s13195-021-00773-z
Identifies pleiotropic loci shared between Alzheimer’s disease and cardiometabolic traits.
2020

Enhanced integrated gradients: improving interpretability of deep learning models using splicing codes as a case study
Genome Biology
·
19 Jun 2020
·
doi:10.1186/s13059-020-02055-7
Improves deep learning interpretability with Enhanced Integrated Gradients using splicing codes as a case study.
2017
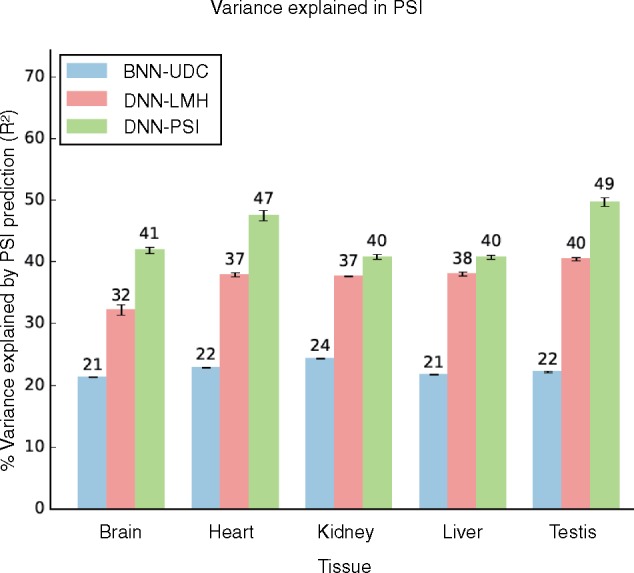
Integrative deep models for alternative splicing
Bioinformatics
·
12 Jul 2017
·
pmc:PMC5870723
Develops integrative deep learning models for predicting alternative splicing regulation.
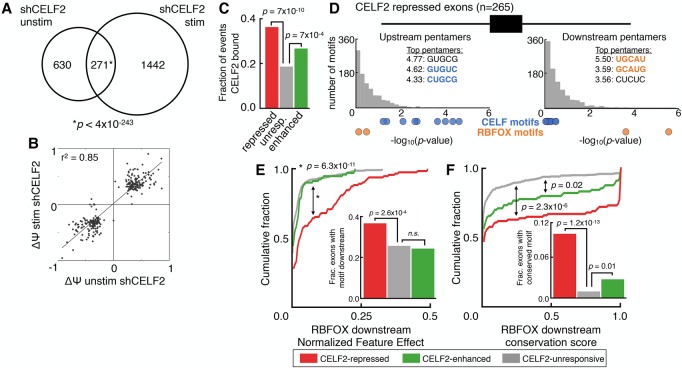
Ancient antagonism between CELF and RBFOX families tunes mRNA splicing outcomes
Genome Research
·
16 May 2017
·
doi:10.1101/gr.220517.117
Reveals how CELF and RBFOX family antagonism shapes mRNA splicing outcomes.
2011

An Optimizing Compiler for Turing Machine Description Language
IUP Journal of Computer Sciences
·
01 Jul 2011
·
iup:tmdl-compiler
Describes an optimizing compiler for a Turing Machine Description Language.